 Print the contig overview to get a clearly arranged version of your contig and its samples. They are shown in the contig overview and mate information for a read can be found in mouse overs.Įditing sample and contig gaps just got easier: You can now move more than one gap at the same time in CodonCode Aligner.Īligner' s contig view allows you to use drag and drop to move the gaps and gives instant visual feedback of the gap movement. Paired ends can be dislpayed for Bowtie2 alignments. In CodonCode Aligner you can now align your sequencing data with Bowtie2. Clicking on a fragment in the gel selects this region in other views and enables copying: Mouse overs display the fragment size as well as start and end base. Gel ladders can be selected, and the gel provides the option to show all fragements in one lane and as separate lanes for each enzyme. Methylation analysis uses raw ABI trace data, which avoids artifacts from trace "normalization" and provides higher accuracy.Īs additional display option for restriction digests, CodonCode Aligner 5.1.1 introduces the virtual gel. Mouse overs also show the calculated methylation ratio, as well as the intensity of the raw peaks. 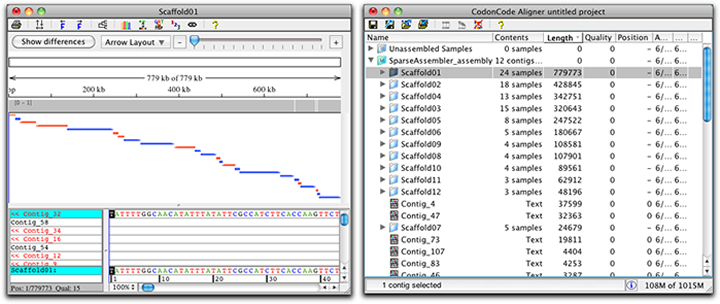
Methylation results are displayed in our features table that can be exported and used to navigate to the methylated base.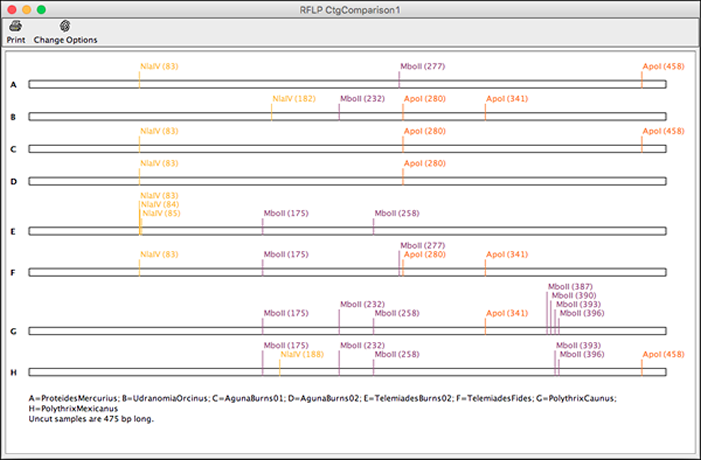 
License keys for version 5.0 will continue to work with version 5.1.ĬodonCode Aligner 5.1 adds functions to analyze methylation of cytosines after bisulfite modification, PCR, and capillary sequencing. Other customers can purchase an upgrade to version 5. Version 5 is a free upgrade for all users with a current update and support agreement as of March 27, 2014. Please note that CodonCode Aligner 5 requires a new license key. CodonCode Aligner 5.1.1 introduced methylation analysis and virtual gels. Additional new features include moving multiple gaps with drag & drop, alignments with Clustal Omega, contig overview printing, and multiple speed and memory improvements for large projects and alignments. This version includes several bug fixes and improved methylation analysis accuracy.ĬodonCode Aligner version 5 added functions to create Bowtie2 alignments and display paired end reads. 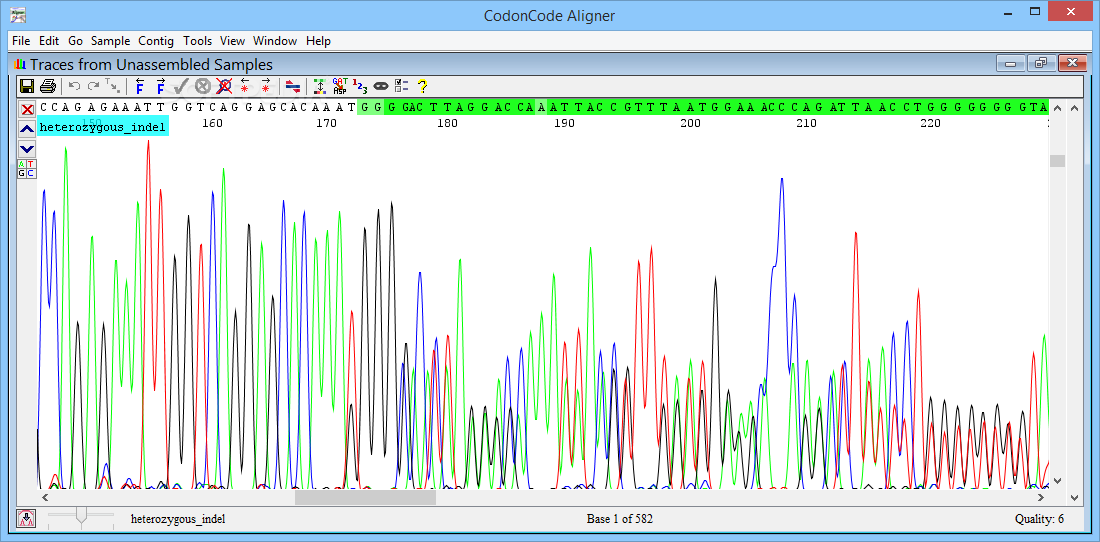
Pricing for Sequencer must be obtained by requesting a quote, which you can do here.The current version of CodonCode Aligner is 5.1.5. It has many new capabilities that can reduce the time required to identify and validate heterozygotes and SNPs in your sequences. Sequencher capabilities include heterozygote and SNP detection and analysis, cDNA to Genomic DNA large gap alignment, comparative sequencing, support for confidence scores, ORF translation, GenBank feature import, and restriction enzyme mapping. Life Science researchers use Sequencher for many diverse DNA sequence analysis applications including de novo gene sequencing, mutation detection, forensic human identification, systematics, and more. First released almost 15 years ago, Sequencher is currently used for sequence analysis tasks in every major genomic and pharmaceutical company as well as numerous academic and government labs in over 40 countries around the world. It works with all automated sequencers and is widely known for its lightning-fast contig assembly, short learning curve, user-friendly editing tools, and superb technical support. Sequencher is the industry standard software for DNA sequence analysis. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
